The BioNLP Open Shared Tasks 2019 workshop co-organized by INRA-MaIAGE and DBCLS (Tokyo) brought together in Hong Kong on November 4 the organizers and participants of the 7 tasks and the scientists in the field.
The tasks cover a wide variety of needs (fine information extraction, ontological standardization, information retrieval, linguistic annotation), languages (English, Spanish) and objects (plants, microorganisms, humans).
The programme, made of 23 presentations and 5 posters, provided an opportunity to present and discuss the tasks and results published by the Association for Computational Linguistics (ACL) in the form of articles selected by an international peer review committee (proceedings available online).
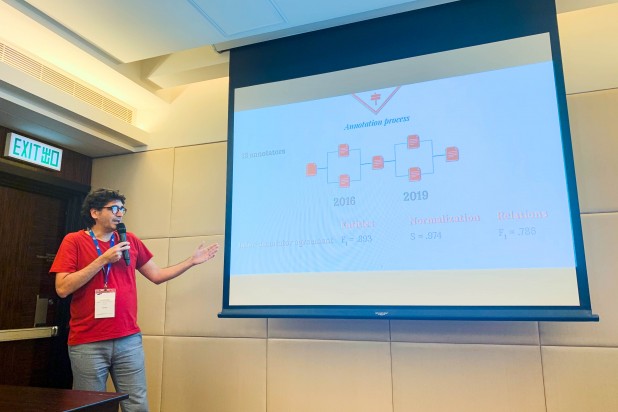
Robert Bossy, research engineer in the Bibliome team (MaIAGE INRA), involved in the evaluation, development, and application of Information Extraction methods in Life Sciences, presented the Bacteria Biotope task at the workshop.
The Bacteria Biotope task in its 4th edition, in addition to the names of microorganisms, habitats and geographical locations, includes phenotypes (e. g. antibiotic resistant) to be extracted from abstracts and articles. The popularity of automatic learning approaches based on neural networks using or not linguistic information is confirmed.
The Bacteria Biotope task presents the challenge of formally representing textual information by a knowledge base through automatic standardization by ontologies.
The participants' systems were able to compensate for the small number of examples of training (small or no data) in terms of the number of classes. All data and results using different metrics can be found on the website and in the task article [Bossy et al., BioNLP-OST 2019].
The best results obtained by participants on the 6 subtasks are several points higher than those of the 2016 edition, despite the increasing complexity, with the exception of the recognition and standardization of microorganism names.
All researchers are encouraged to continue to use in the future the data from the Bacteria Biotope task, the evaluation service and supporting resources (e. g. word embeddings), both public and online. A future call for articles in a special issue of the newspaper will extend their exploitation as in previous editions.
References :
Robert Bossy, Louise Deléger, Estelle Chaix, Mouhamadou Ba, and Claire Nédellec. 2019. Bacteria Biotope at BioNLP Open Shared Tasks 2019. In Proceedings of the BioNLP Open Shared Task 2019 Workshop joint to EMNLP-IJCNLP 2019, Association for Computational Linguistics pub., pages 121-131, Hong Kong 4 novembre 2019.
Kim Jin-Dong, Nédellec Claire, Bossy Robert, Deléger Louise (éditeurs). Proceedings of The 5th Workshop on BioNLP Open Shared Tasks joint to EMNLP-IJCNLP 2019, Association for Computational Linguistics pub., Hong Kong 4 novembre 2019.